Please note: The web browser must be configured to accept cookies. Both of Java and JavaScript must be enabled. Java JRE/JDK must be newer than version 1.4.2.
Virus Human Interactome Network Map
Applying a systematic integrated pipeline 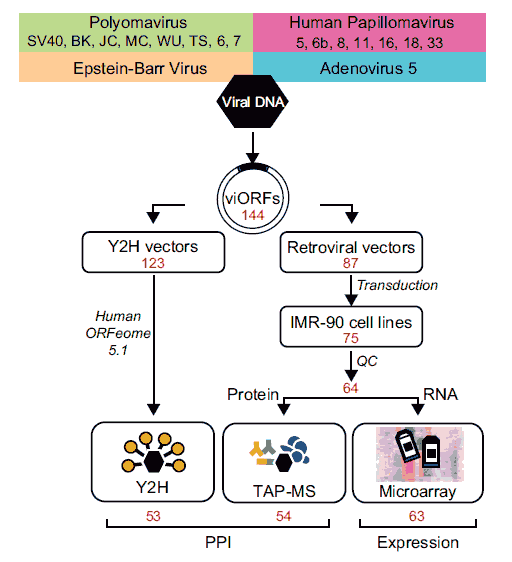 we investigated at genome-scale (Rozenblatt-Rosen et al, Nature 487:491-5) perturbations of host interactome and transcriptome networks induced by individual gene products encoded by members of four functionally related, yet biologically distinct, families of DNA tumor viruses: several polyomaviruses (PyV), simian virus 40 (SV40) in particular; six classes of human papillomaviruses (HPV), high-risk (for carcinogenesis) mucosal HPV16 and HPV18, low risk mucosal HPV6b and HPV11, and cutaneous HPV5 and HPV8; Epstein-Barr Virus (EBV); and Adenovirus 5 (Ad5).
we investigated at genome-scale (Rozenblatt-Rosen et al, Nature 487:491-5) perturbations of host interactome and transcriptome networks induced by individual gene products encoded by members of four functionally related, yet biologically distinct, families of DNA tumor viruses: several polyomaviruses (PyV), simian virus 40 (SV40) in particular; six classes of human papillomaviruses (HPV), high-risk (for carcinogenesis) mucosal HPV16 and HPV18, low risk mucosal HPV6b and HPV11, and cutaneous HPV5 and HPV8; Epstein-Barr Virus (EBV); and Adenovirus 5 (Ad5).
Binary Interactome Network
To map binary physical interactions at genome-scale between viral and host proteins we used a stringent implementation of the yeast two-hybrid (Y2H) system, We ultimately screened 123 viral open reading frames (viORFs) against a collection of ~13,000 human ORFs. The viral-host interactome so obtained contained 454 validated binary interactions between 53 viral proteins and 307 human target proteins (search the interactome data).
Co-complex Interactome Network
To map co-complex associations at proteome-scale between viral proteins and the host proteome, we implemented a tandem affinity purification protocol followed by mass spectrometry (TAP-MS). For TAP-MS we generated expression constructs of each viral ORF fused to a tandem epitope tag, and introduced each expression construct into IMR90 normal human diploid fibroblast cells. The intersection of two independent TAP-MS experiments yielded 3,787 reproducibly mapped viral-host co-complex associations involving 54 viral proteins and the products of 1,079 unambiguously identified host genes (search the interactome data).
Transcriptome Profiling Data
To characterize virally induced transcriptional perturbations at genome-scale, we completed microarray analyses on the TAP-tagged viral ORF-expressing IMR90 cell lines, scoring significant host gene expression changes. To identify patterns of host transcriptional perturbation common across the set of viral proteins, model-based clustering was used to construct clusters from the most frequently perturbed host genes. We identified 31 clusters of five or more genes, most of them enriched for GO terms and KEGG pathways (download the transcriptome dataset).
VirHost set of potential cancer genes
The viral target proteins, identified through binary interaction, co-complex
associations, and transcription factor binding site analyses, preferentially targeted host proteins altered in cancer. For optimization of the stringency of potential cancer enrichment analyses, the TAP-MS co-complex viral targets were restricted to those identified by three or more unique peptides, a choice corresponding to an experimental reproducibility rate greater than 90%. In the resulting stringent candidate set of 947 host target genes (the “VirHost” set) tumor suppressor genes were significantly overrepresented. (download the VirHost dataset).