Please note: The web browser must be configured to accept cookies. Both of Java and JavaScript must be enabled. Java JRE/JDK must be newer than version 1.4.2.
Human Interactome Project
Please visit our new web portal site: Human Reference Protein Interactome
One of the long-term goals of CCSB is to generate a first reference map of the human protein-protein interactome network. To reach this target, we are mapping binary protein-protein interactions by systematically interrogating all pairwise combinations of predicted gene products in defined search spaces using proteome-scale technologies. Our approach is to map high-quality binary protein-protein interactions using a primary yeast two-hybrid assay (Y2H) followed by orthogonal validation by alternative binary assays. Currently, we have completed two proteome-scale efforts (HI-I-05 and HI-II-14) based on the Gateway-cloned ORF collections available at the time of these experiments (http://horfdb.dfci.harvard.edu). Other efforts to optimize our pipeline and benchmark the quality of our maps have also led to the production of systematic maps although of relatively smaller size (Venkatesan-09 and Yu-11). In 2013, we have extracted, filtered, and benchmarked interaction data from 7 public databases to extract a high-quality binary literature dataset (Lit-BM-13) comprising all protein-protein interactions that are binary and supported by at least two traceable pieces of evidence (publications and/or methods).
All individual datasets are described in further details here below and are freely available to the research community through the search engine or via download. Preliminary data from ongoing projects are also described but are only available to registered users for download here.
Proteome-scale efforts
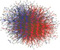 HI-I-05: Our first iteration at mapping the human interactome (Rual et al Nature 2005) screened a space (Space I) of ~8,000 ORFs corresponding to ~7,000 genes, and identified ~2,700 high-quality binary interactions. This search space represents ~12% of the complete search space, assuming a total of ~20,000 protein-coding genes and limiting the scope to one variant per gene. This dataset is freely available to the research community through the search engine or via download.
HI-I-05: Our first iteration at mapping the human interactome (Rual et al Nature 2005) screened a space (Space I) of ~8,000 ORFs corresponding to ~7,000 genes, and identified ~2,700 high-quality binary interactions. This search space represents ~12% of the complete search space, assuming a total of ~20,000 protein-coding genes and limiting the scope to one variant per gene. This dataset is freely available to the research community through the search engine or via download.
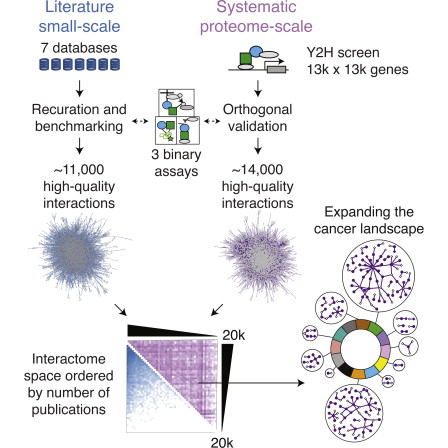 HI-II-14: The second phase of the human interactome project (Rolland et al Cell 2014) generated a dataset of ~14,000 binary interactions following two screens of a matrix of ~13,000 x 13,000 proteins (Space II). This search space covers ~42% of the complete search space, a more than 3 fold increase with respect to our first attempt. This dataset is freely available to the research community through the search engine or via download.
HI-II-14: The second phase of the human interactome project (Rolland et al Cell 2014) generated a dataset of ~14,000 binary interactions following two screens of a matrix of ~13,000 x 13,000 proteins (Space II). This search space covers ~42% of the complete search space, a more than 3 fold increase with respect to our first attempt. This dataset is freely available to the research community through the search engine or via download.
HI-III: The next phase of the human interactome project is underway. The human ORF collection being screened has been expanded to ~17,500 unique genes (Space III) and covers ~77% of the complete search space. The list of genes being screened is provided here. Preliminary data produced by this ongoing effort are provided prepublication for registered users here.
Other datasets
 Venkatesan-09: To estimate the coverage and size of the human interactome (Venkatesan et al Nat Methods 2009), four Y2H screens were performed on a set of ~1,800 DB-X fusion proteins (or baits, representing ~1,700 unique genes) against ~1,800 AD-Y proteins (or preys, representing ~1,800 unique genes), corresponding to ~10% of the available genes and ~1% of the full search space. This dataset contains ~200 high-quality Y2H interactions, and is freely available to the research community through the search engine or via download.
Venkatesan-09: To estimate the coverage and size of the human interactome (Venkatesan et al Nat Methods 2009), four Y2H screens were performed on a set of ~1,800 DB-X fusion proteins (or baits, representing ~1,700 unique genes) against ~1,800 AD-Y proteins (or preys, representing ~1,800 unique genes), corresponding to ~10% of the available genes and ~1% of the full search space. This dataset contains ~200 high-quality Y2H interactions, and is freely available to the research community through the search engine or via download.
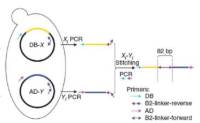 Yu-11: To develop a novel Stitch-seq interactome mapping protocol, a Y2H screen was carried out inside Space II (Yu et al Nat Methods 2011). Stitch-seq combines PCR stitching with next-generation sequencing, and increases the efficiency and cost effectiveness of Y2H screening. The resulting dataset contains ~1,200 interactions among proteins encoded by ~1,100 human genes. This dataset is freely available to the research community through the search engine or via download.
Yu-11: To develop a novel Stitch-seq interactome mapping protocol, a Y2H screen was carried out inside Space II (Yu et al Nat Methods 2011). Stitch-seq combines PCR stitching with next-generation sequencing, and increases the efficiency and cost effectiveness of Y2H screening. The resulting dataset contains ~1,200 interactions among proteins encoded by ~1,100 human genes. This dataset is freely available to the research community through the search engine or via download.
Test Space: To develop, optimize, and benchmark improvements to the mapping pipeline used for ongoing efforts (HI-III), twelve independent, reciprocal screens on a search space of ~1,800 x ~1,800 genes were completed, constituting ~1% of the full search space. Preliminary data produced by this ongoing effort are provided prepublication for registered users here.
High-quality non-systematic Literature dataset
Lit-BM-13: In 2013, we extracted interaction data from BIND, BioGRID, DIP, HPRD, MINT, IntAct, and PDB to generate a high-quality binary literature dataset comprising ~11,000 protein-protein interactions that are binary and supported by at least two traceable pieces of evidence (publications and/or methods) (Rolland et al Cell 2014). Although this dataset does not result from a systematic investigation of the interactome search space and should thus be used with caution for any network topology analyses, it represents valuable interactions for targeted studies and is freely available to the research community through the search engine or via download.