Please note: The web browser must be configured to accept cookies. Both of Java and JavaScript must be enabled. Java JRE/JDK must be newer than version 1.4.2.
Yeast Interactome Project
The interactome of an organism is the network formed by the full set of physical interactions that can occur in a physiologically relevant dynamic range between all its macromolecules, including protein-protein, DNA-protein, and RNA-protein interactions. Our goal is to develop a more complete yeast protein-protein interaction (PPI) map, leveraging the knowledge we’ve gained from systematic binary interaction mapping efforts. Yeast affords us the opportunity to interrogate a comprehensive set of protein-coding genes and to integrate our systematic interaction data with high quality literature-curated information and with other systematic datasets that are amenable to orthogonal validation. The generation of interactome network maps with increasing sensitivity and quality is a necessary, albeit not sufficient, aspect in the quest of generating predictive macromolecular models at the scale of the whole cell. We developed a conceptual framework to assess interactome models based on quantitative benchmarking of screening and validation assays against reference sets. With this we have shown that the set of high quality, systematic yeast interactome data sets cover ~10% to ~20% of the predicted yeast binary interactome, albeit interrogating less than 75% of yeast protein-coding genes. Utilizing novel technologies we aim to extend to 50% to enable a profoundly improved understanding of the interactome network of budding yeast. For quality control, binary interactions are validated in standardized orthogonal interaction assays, allowing us to thus generate and analyze a third generation, high-quality binary interactome map of S. cerevisiae.
Previous efforts
YI-I: We carried out a comparative quality assessment of current yeast interactome datasets, demonstrating that high-throughput yeast two-hybrid (Y2H) provides high-quality binary interaction information. As a large fraction of the yeast binary interactome remains to be mapped, we developed an empirically-controlled mapping framework to produce a "second-generation" high-quality, high-throughput Y2H dataset covering ~20% of all yeast binary interactions. Both Y2H and affinity purification followed by mass spectrometry (AP/MS) data are of equally high-quality but of a fundamentally different and complementary nature resulting in networks with different topological and biological properties. This binary map is enriched for transient signaling interactions and inter-complex connections with a highly significant clustering between essential proteins. Rather than correlating with essentiality, protein connectivity correlates with genetic pleiotropy.
"High-Quality Binary Protein Interaction Map of the Yeast Interactome Network" Yu et al Science 2008.
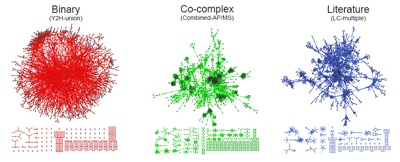
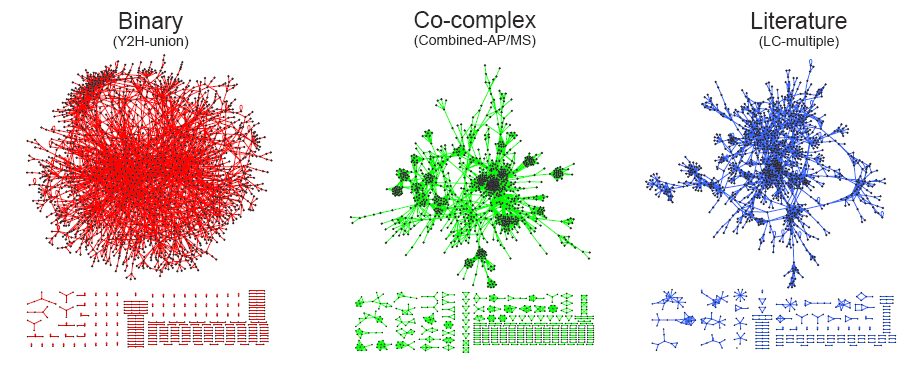
Graphical representation of three different types of yeast interactome datasets. The structure of the binary interactome network is obviously different from the structure of the co-complex interactome network. The network structure of the literature-curated dataset resembles that of the co-complex dataset, even though the literature-curated datasets are reported to contain mostly binary interactions. All datasets can be downloaded here.